Total RNA Extraction Kit (Plant)
No:DMRPL100 & 300
For research use only
Sample: 100 mg of fresh plant tissue or 25 mg of dry plant tissue
Yield : up to 30μg
Introduction
The BioDiamond Total RNA Extraction Kit provides a fast, simple, and cost-effective method for isolation of total RNA from plant samples. Detergents and chaotropic salt are used to lyse cells and inactivate RNase. The specialized high-salt buffering system allows RNA species longer than 100 bases to bind to the the glass fiber matrix of the spin column . The BioDiamond Total RNA Extraction Kit is suitable for a variety of routine applications including RT-PCR, cDNA Synthesis, Northern Blotting, Differential display, Primer Extension and mRNA Selection. The entire procedure can be completed within 60 minutes.
Kit Contents
NOTE
★ Always wear latex or vinyl gloves while handling reagents and RNA samples to prevent RNase contamination.
★ Add ethanol (96–100%) to Buffer W2, shake before use (see bottle label for volume).
★ Check Buffers before use for salt precipitation. Redissolve any precipitate by warming to 37°C.
★ Buffers RP and W1 contain irritants. Wear gloves when handling these buffers.
Quality Control
In accordance with FairBiotech’s ISO-certified Total Quality Management System, the quality of the BioDiamond Total RNA Extraction Kit is tested on a lot-to-lot basis to ensure consistent product quality.
Additional requirements
*Ethanol (96~100%)
*Isopropanol
*RNase-free pipet tips and 1.5 ml microcentrifuge tubes
*ß-mercaptoethanolc
#For Optional Step (DNA Residue Degradation):
Add 2 µl DNAse I (2KU/ml) and 10 µl reaction buffer {300 mM Tris-HCl (pH 7.5), 60 mM MnCl2, 300 µg/ml BSA } to the 50μl final product. Let stand for 10 minutes at room temperature (at 25ºC).
Protocol
Sample Preparation
- Cut off 100 mg of fresh plant tissue or 50 mg of dry plant tissue.
- Grind the sample under liquid nitrogen to a fine powder using a mortar and pestle.
- Add 1 ml of RP Buffer and 10 μl of ß-mercaptoethanol to the sample in the mortar and grind the sample until it is completely dissolved.
- Transfer the dissolved sample to a RNase-free 1.5 ml microcentrifuge tube. Incubate at 70ºC for 30 minutes. (invert the tube every 10 minutes)
- Centrifuge at 2-8ºC at 14-16,000 x g for 10 minutes. Transfer the supernatant to a new 1.5 ml microcentrifuge tube.
- Add a ½ volume of Isopropanol to the sample from Step 1 and shake vigorously (e.g. add 250 µl of Isopropanol to 500 µl of sample).
- Place a RP Column in a 2 ml Collection Tube.
- Transfer the sample mixture to the RP Column. Centrifuge at 14-16,000 x g for 30 seconds.
- Discard the flow-through and transfer the remaining mixture to the same RP Column.
- Centrifuge at 14-16,000 x g for 30 seconds.
- Discard the flow-through and place the DP Column back in the 2 ml Collection Tube.
- Add 400 µl of W1 Buffer to the RP Column. Centrifuge at 14-16,000 x g for 30 seconds.
- Discard the flow-through and place the RP Column back in the 2 ml Collection Tube.
- Add 600 µl of W2 Buffer (ethanol added) into the RP Column. Centrifuge at 14-16,000 x g for 30 seconds.
- Discard the flow-through and place the RP Column back in the 2 ml Collection Tube.
- Centrifuge again for 3 minutes at 14-16,000 x g to dry the column matrix.
- To elute RNA, place the RP Column in a new RNase-free 1.5 ml microcentrifuge tube.
- Add 50-200μl REL Buffer to the center of each RP Column, let stand for 2 minutes, and centrifuge at 14-16,000 x g for 2 minutes.
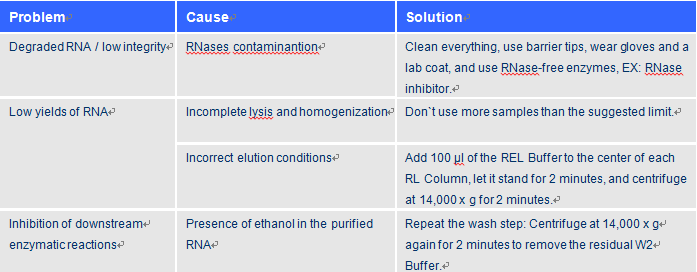
Troubleshooting
| dmrpl100___300.pdf | |
| File Size: | 242 kb |
| File Type: | |