PCR Clean Up/Gel Extraction Kit
No:DMDSPG100 & 300
For research use only
Sample: up to 100μl of PCR Product or 300 mg of Agarose Gel
Recovery: up to 95%
Introduction
The BioDiamond PCR Clean Up/Gel Extraction Kit provides a cost-effective system for fast and easy isolation of DNA fragments from PCR reactions, agarose gels, or enzymatic reactions. DNA fragments (50bp-5Kb) in specialized buffers are bound by the glass fiber matrix of the spin column (1, 2) while contaminants pass through the column. Impurities are efficiently washed away, and pure DNA is eluted with Tris buffer or water without phenol extraction or alcohol precipitation. DNA purified with the kits is suitable for any subsequent application, such as ligation and transformation, sequencing, restriction enzyme digestion, labeling, PCR, in vitro transcription, or microinjection. The entire procedure can be completed within 15-20 minutes.
Kit Contents
NOTE
★ Add ethanol (96–100%) to Buffer W2, shake before use (see bottle label for volume).
★ Check Buffers before use for salt precipitation. Redissolve any precipitate by warming to 37°C.
★ Buffers DE and W1 contain irritants. Wear gloves when handling these buffers.
Quality Control
In accordance with FairBiotech’s ISO- certified Total Quality Management System, the quality of the BioDiamond PCR Clean Up /Gel Extraction Kit is tested on a lot to lot basis to ensure consistent product quality.
Additional requirements
*Ethanol (96~100%) *1.5 ml microcentrifuge tubes
Protocol
Step 1 Sample Preparation
Gel Extraction
- Excise the DNA fragment from the agarose gel. Transfer up to 300 mg of the gel slice to a 1.5 ml microcentrifuge tube. Add 500 μl Buffer DE to the sample and mix by vortex.
- Incubate at 60°C for 10 minutes (or until the gel slice has completely dissolved). During the incubation, mix by vortexing the tube every 2–3 minutes. Cool the dissolved sample mixture to room temperature.
- Add 500 μl Buffer DE to 100 μl of the PCR reaction and mix by vortex.
- Place a SDE Column in a Collection Tube. Apply the supernatant (from step 1) to the DE column by decanting or pipetting.
- Centrifuge at 14,000 x g for 30 seconds. Discard the flow-through and place the DE column back into the same Collection tube. (The maximum volume of the DE column reservoir is 800 μl. If the sample mixture is more than 800 µl, repeat the DNA Binding Step)
- Add 400 µl of Buffer W1 into the SDE Column. Centrifuge at 14,000 x g for 30 seconds. Discard the flow-through and place the SDE column back into the same Collection tube.
- Add 600 µl of Buffer W2 (ethanol added) into the SDE Column. Centrifuge at 14,000 x g for 30 seconds. Discard the flow-through and place the SDE column back into the same Collection tube.
- Centrifuge at 14,000 x g again for 2 minutes to remove residual Buffer W2.
- To elute DNA, place the SDE column in a clean 1.5 ml microcentrifuge tube.
- Add 50-200 μl Buffer EL or water (pH is between 7.0 and 8.5) to the center of eachS DE column, let stand for 2 min, and centrifuge at 14,000 x g for 2 min.
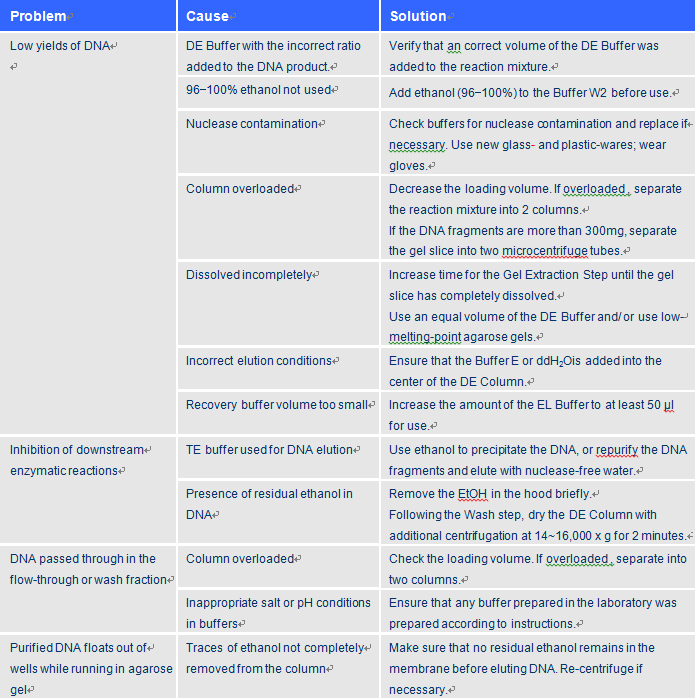
Troubleshooting
| dmdspg100___300.pdf | |
| File Size: | 174 kb |
| File Type: | |