Genomic DNA Extraction Kit (Blood/Cultured Cell/Bacteria)
No:DMGBA100 & 300
For research use only
Sample: up to 300 μl of whole blood, 200 μl of buffy coat, 107mammalian cells, 5×107 fungus cells and 109 bacterial cells
Yield : up to 50 μg
Introduction
This BioDiamond Genomic DNA Extraction Kit (Cultured Cell/Blood) was designed specifically for genomic DNA isolation from whole blood, frozen blood, buffy coat, cultured animal/bacterial cells and fungal cells. Its unique buffer system ensures genomic DNA with high yield and good quality from samples while the spin column purifies and concentrates genomic DNA products previously isolated with the buffer system. The entire procedure can be completed in 1 hour without phenol/chloroform extraction needs. Purified genomic DNA is suitable for use in PCR or other enzymatic reactions.
Kit Contents
NOTE
★ Add ethanol (96–100%) to Buffer W2, shake before use (see bottle label for volume).
★ Check Buffers before use for salt precipitation. Redissolve any precipitate by warming to 37°C.
★ Buffers W1 contain irritants. Wear gloves when handling these buffers.
Quality Control
In accordance with FairBiotech’s ISO-certified Total Quality Management System, the quality of the BiodIAMOND Genomic DNA Extraction Kit (Cultured cell/ Blood) tested on a lot-to-lot basis to ensure consistent product quality.
Additional requirements
*microcentrifuge tubes
* absolute ethanol
*RNase A (10 mg/ml)
* For Gram-positive bacteria samples: lysozyme buffer (20 mg/ml lysozyme; 20 mM Tris-HCl; 2 mM EDTA; 1% TritonX-100; pH 8.0, prepare the lysozyme buffer immediately prior to use)
* For Fungus samples: lyticase or zymolase, sorbitol buffer (1.2 M sorbitol;10 mM CaCl2; 0.1 M Tris-HCl pH 7.5; 35 mM mercaptoethanol)
Protocol
Step 1 Sample Cells Harvesting
Fresh whole Blood or Buffy Coat
- Collect blood in EDTA-Na2 treated collection tubes (or other anticoagulant mixtures).
- Transfer up to 300 μl of blood or 200 μl of buffy coat to a sterile1.5 ml microcentrifuge tube.
- Add 900 μl of CR Buffer and mix by inversion.
- Incubate the tube at room temperature for 10 minutes (invert twice during incubation).
- Centrifuge for 5 minutes at 4,000 x g. Remove the supernatant completely and resuspend the cells in 50 µl of CR Buffer by pipetting the pellet up and down.
- Transfer cultured mammalian cells (up to 107) to a sterile 1.5 ml microcentrifuge tube.
- Centrifuge at 6,000 x g for 1 minute. Remove the supernatant completely and resuspend the cells in 50 µl of CR Buffer by pipetting the pellet up and down.
- Transfer cultured bacterial cells (up to 109) to a sterile 1.5 ml microcentrifuge tube.
- Centrifuge at 12,000 x g for 1 minute. Remove the supernatant completely and resuspend the cells in 50 µl of CR Buffer by pipetting the pellet up and down.
- Transfer cultured bacterial cells (up to 109) to a sterile 1.5 ml microcentrifuge tube.
- Centrifuge at 12,000 x g for 1 minute. Remove the supernatant completely and resuspend the cells in 100 µl of lysozyme Buffer by pipetting the pellet up and down. Incubate at 37ºC for 30 minutes.
- Transfer fungus cells (up to 108) to a sterile 1.5 ml microcentrifuge tube.
- Centrifuge at 6,000 x g for 5 minute. Remove the supernatant completely and resuspend the cells in 600 µl of sorbitol Buffer by pipetting the pellet up and down.
- Add 200 U of lyticase or zymolase. Incubate at 30ºC for 30 minutes.
- Centrifuge the mixture for 10 minutes at 2,000 x g to harvest the spheroplast. Remove the supernatant completely and resuspend the cells in 50 µl of CR Buffer by pipetting the pellet up and down.
- Add 300 µl of CC Buffer to the resuspended cells from Step 1 and mix by vortex.
- Incubate at 60ºC for 10 minutes or until the sample lysate is clear. During incubation, invert the tube every 3 minutes. #Pre-heat the Elution Buffer to 60ºC for Step 6 DNA Elution.
RNA Degradation (If RNA-free genomic DNA is required, perform this optional step.)
- Add 5 µl of RNase A (10 mg/ml) to the sample lysate and mix by vortex. Incubate at room temperature for 5 minutes.
- Add 400 µl of CB Buffer to the sample from Step 2 and shake vigorously.
- Centrifuge at 12,000 x g for 1 minute. (Do not go over 1 minute)
- Place a CC Column in a 2 ml Collection Tube.
- Transfer the clear supernatant completely from the previous step to the CC Column.
- Centrifuge at 14,000 x g for 30 seconds.
- Discard the flow-through and place the CC Column back in the 2 ml Collection Tube.
- Add 400 µl of W1 Buffer into the CC Column. Centrifuge at 14,000 x g for 30 seconds.
- Discard the flow-through and place the CC Column back into the same Collection tube.
- Add 600 µl of W2 Buffer (Ethanol added) into the CC Column. Centrifuge at 14,000 x g for 30 seconds.
- Discard the flow-through and place the CC Column back into the same Collection tube.
- Centrifuge at 14,000 x g again for 2 minutes to remove residual W2 Buffer.
- Transfer the dried CC Column to a new 1.5 ml microcentrifuge tube.
- Add 50-200 µl of Pre-Heated EL Buffer or TE into the center of the column matrix.
- Let stand at 60ºC for 3 minutes.
- Centrifuge for 2 minutes at 14,000 x g to elute the purified DNA.
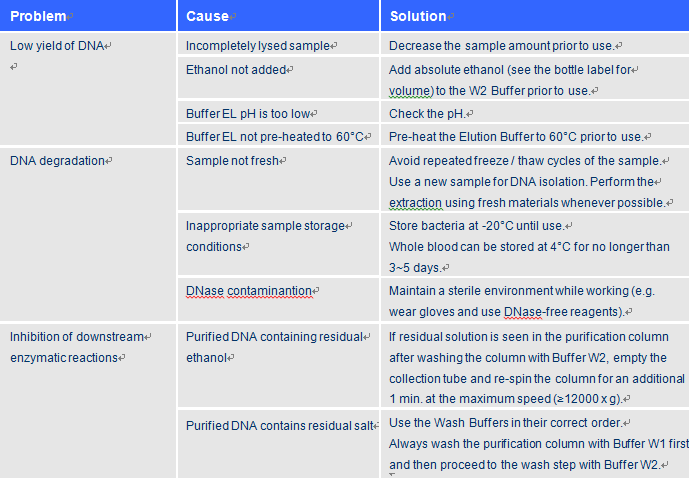
Troubleshooting
| dmgba100___300.pdf | |
| File Size: | 268 kb |
| File Type: | |